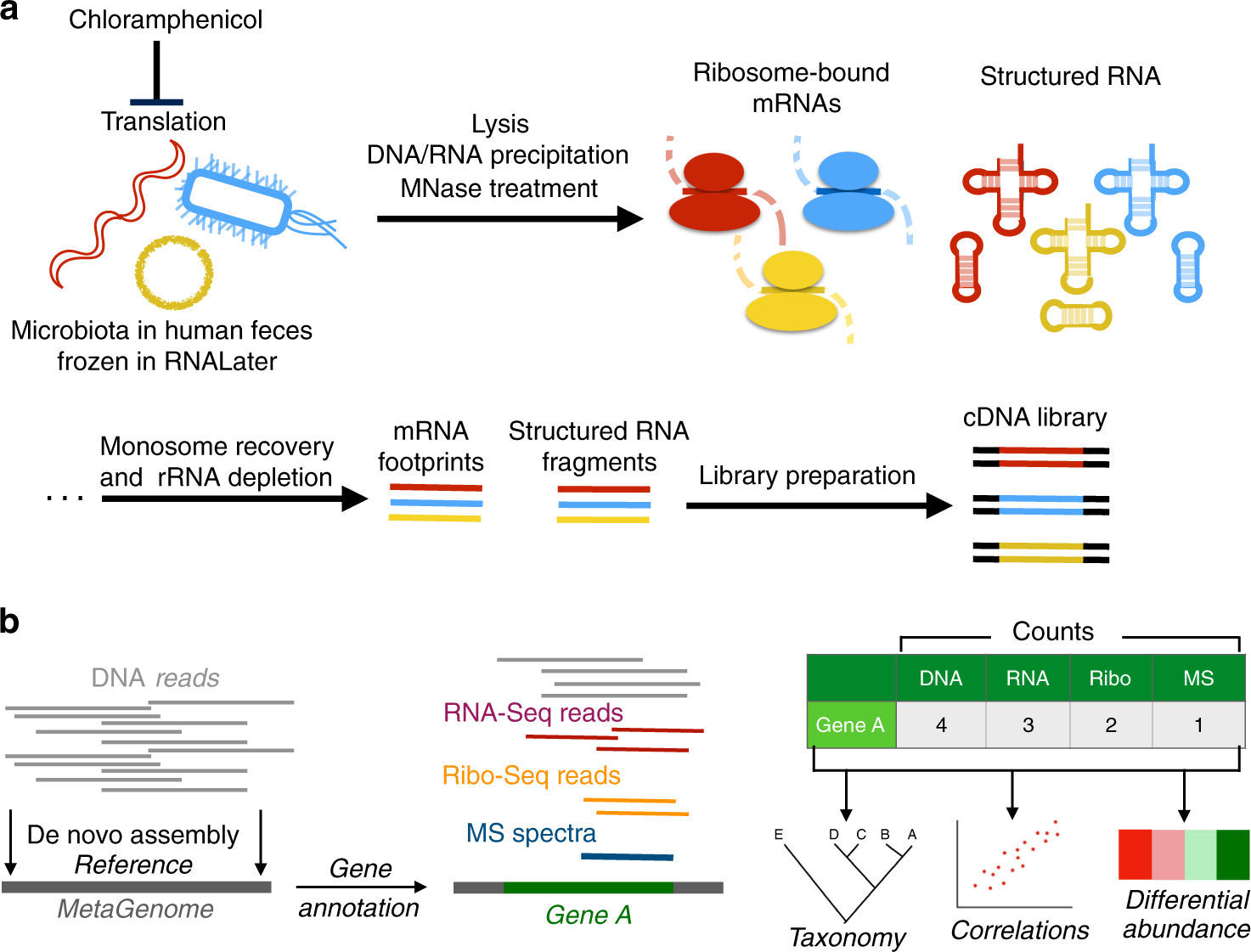
Translational changes induced by acute sleep deprivation uncovered by TRAP- Seq | Molecular Brain | Full Text
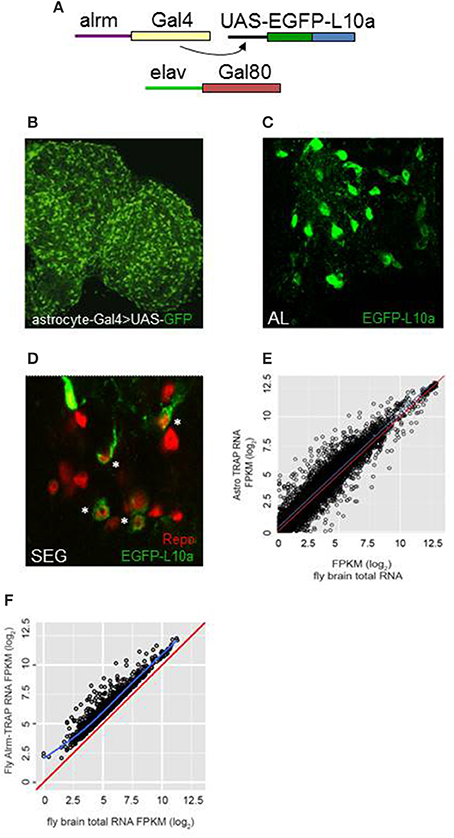
Frontiers | TRAP-seq Profiling and RNAi-Based Genetic Screens Identify Conserved Glial Genes Required for Adult Drosophila Behavior

TrapSeq: An RNA Sequencing-Based Pipeline for the Identification of Gene- Trap Insertions in Mammalian Cells - ScienceDirect

TRAP-Seq Identifies Differentially Translating mRNAs in Fmr1 –/y CA1... | Download Scientific Diagram

Overview of TRAP-Seq experiment and qPCR validation. (A) Schematics... | Download Scientific Diagram
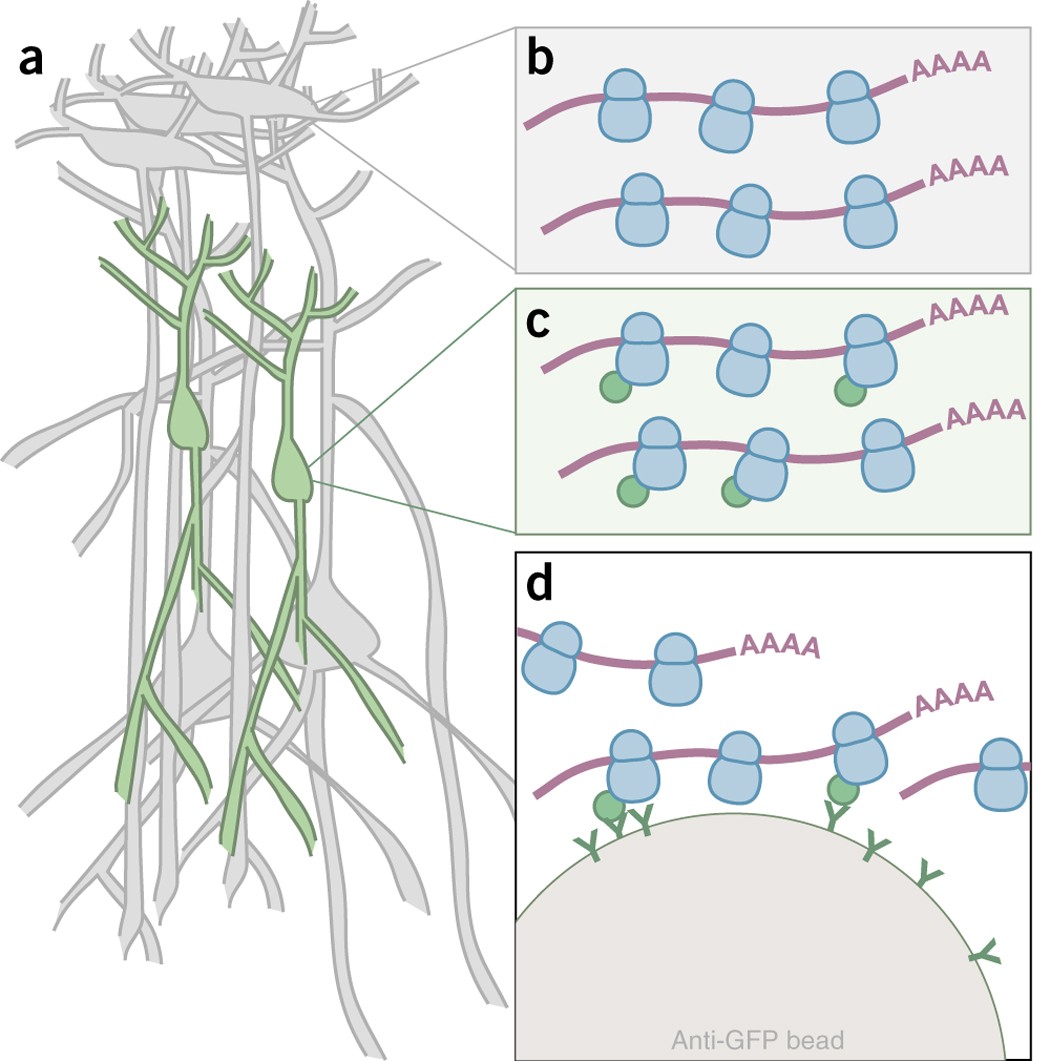
Cell type–specific mRNA purification by translating ribosome affinity purification (TRAP) | Nature Protocols

Translating Ribosome Affinity Purification (TRAP) Followed by RNA Sequencing Technology (TRAP-SEQ) for Quantitative Assessment of Plant Translatomes | SpringerLink

Translational changes induced by acute sleep deprivation uncovered by TRAP- Seq | Molecular Brain | Full Text

TrapSeq: An RNA Sequencing-Based Pipeline for the Identification of Gene- Trap Insertions in Mammalian Cells - ScienceDirect

TRAP-SEQ – Translating Ribosome Affinity Purification (TRAP) Followed by RNA Sequencing Technology | RNA-Seq Blog
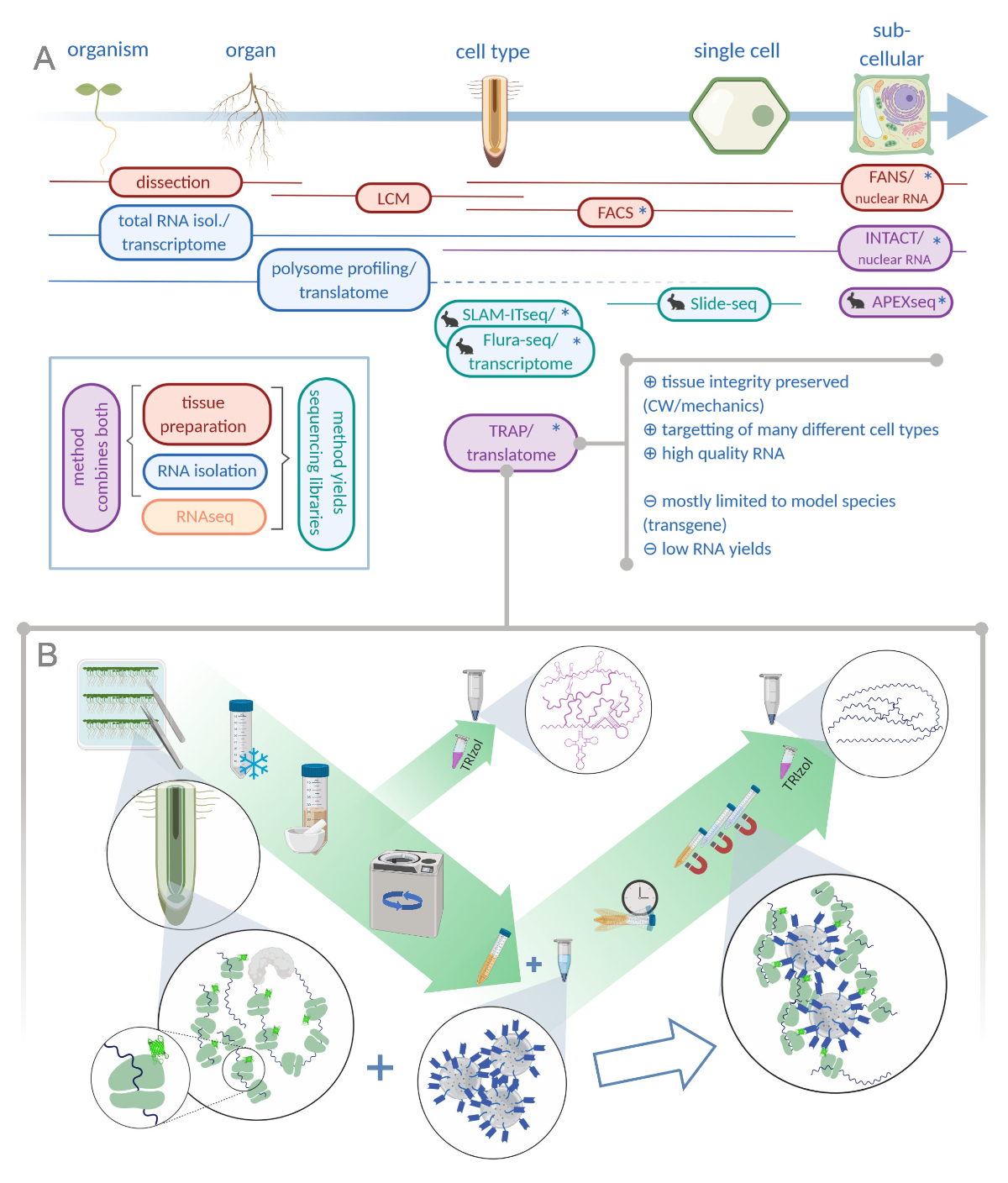
Translating Ribosome Affinity Purification (TRAP) to Investigate Arabidopsis thaliana Root Development at a Cell Type-Specific Scale | Protocol

Rapid Changes in the Translatome during the Conversion of Growth Cones to Synaptic Terminals - ScienceDirect

Quantification of mRNA ribosomal engagement in human neurons using parallel translating ribosome affinity purification (TRAP) and RNA sequencing - ScienceDirect

Validation of the applicability of TRAP-seq to study organ-specific... | Download Scientific Diagram

Evaluation of TRAP-seq by molecular profiling of the Emx1-lineage in... | Download Scientific Diagram

Translational changes induced by acute sleep deprivation uncovered by TRAP- Seq | Molecular Brain | Full Text