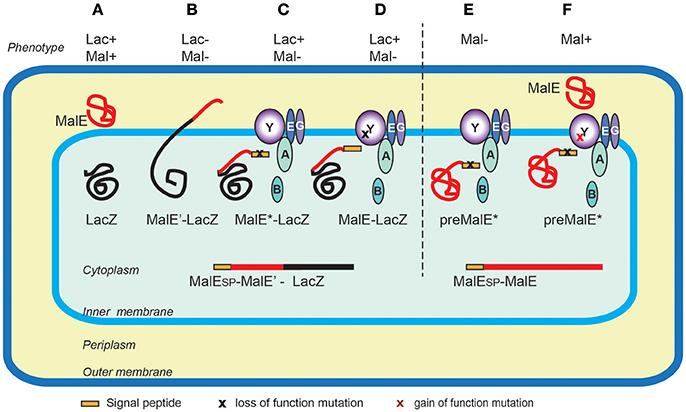
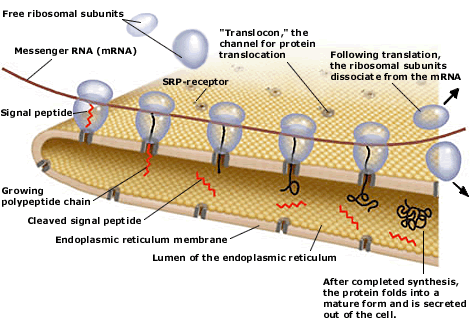
Design of an improved universal signal peptide based on the α-factor mating secretion signal for enzyme production in yeast | Centro de Investigaciones Biológicas Margarita Salas - CIB Margarita Salas

Signal Peptide Efficiency: From High-Throughput Data to Prediction and Explanation | ACS Synthetic Biology

Differential role of segments of α-mating factor secretion signal in Pichia pastoris towards granulocyte colony-stimulating factor emerging from a wild type or codon optimized copy of the gene | Microbial Cell Factories

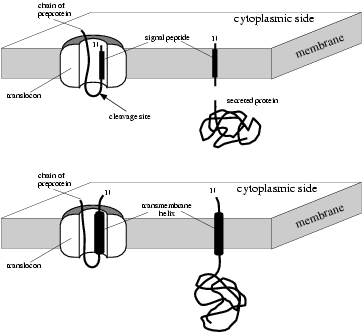
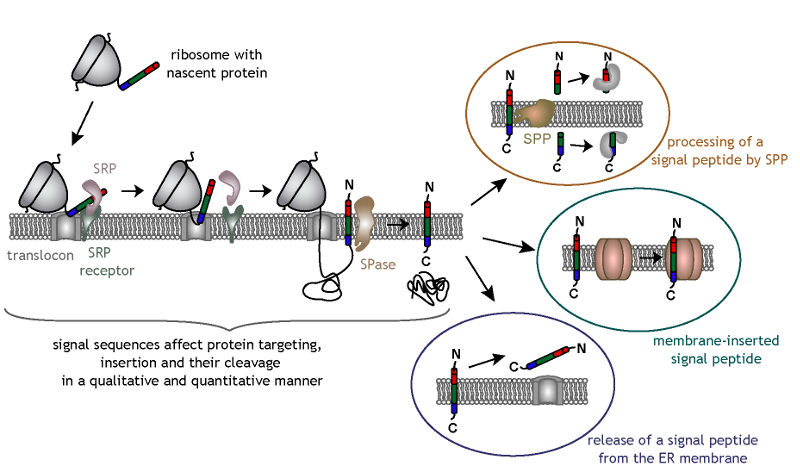
Signal peptide substrates of different classes of signal peptidases.... | Download Scientific Diagram

Signal peptide cleavage of precursor proteins by signal peptidases. Sig- | Download Scientific Diagram
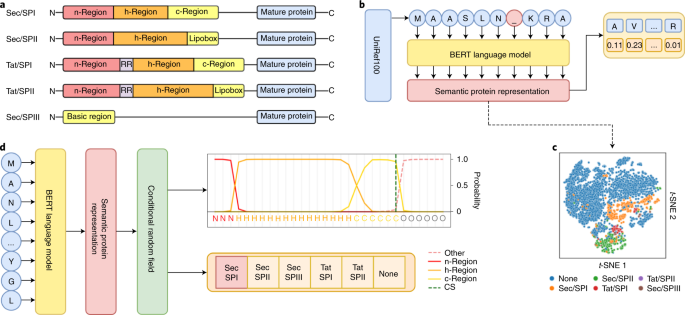
SignalP 6.0 predicts all five types of signal peptides using protein language models | Nature Biotechnology
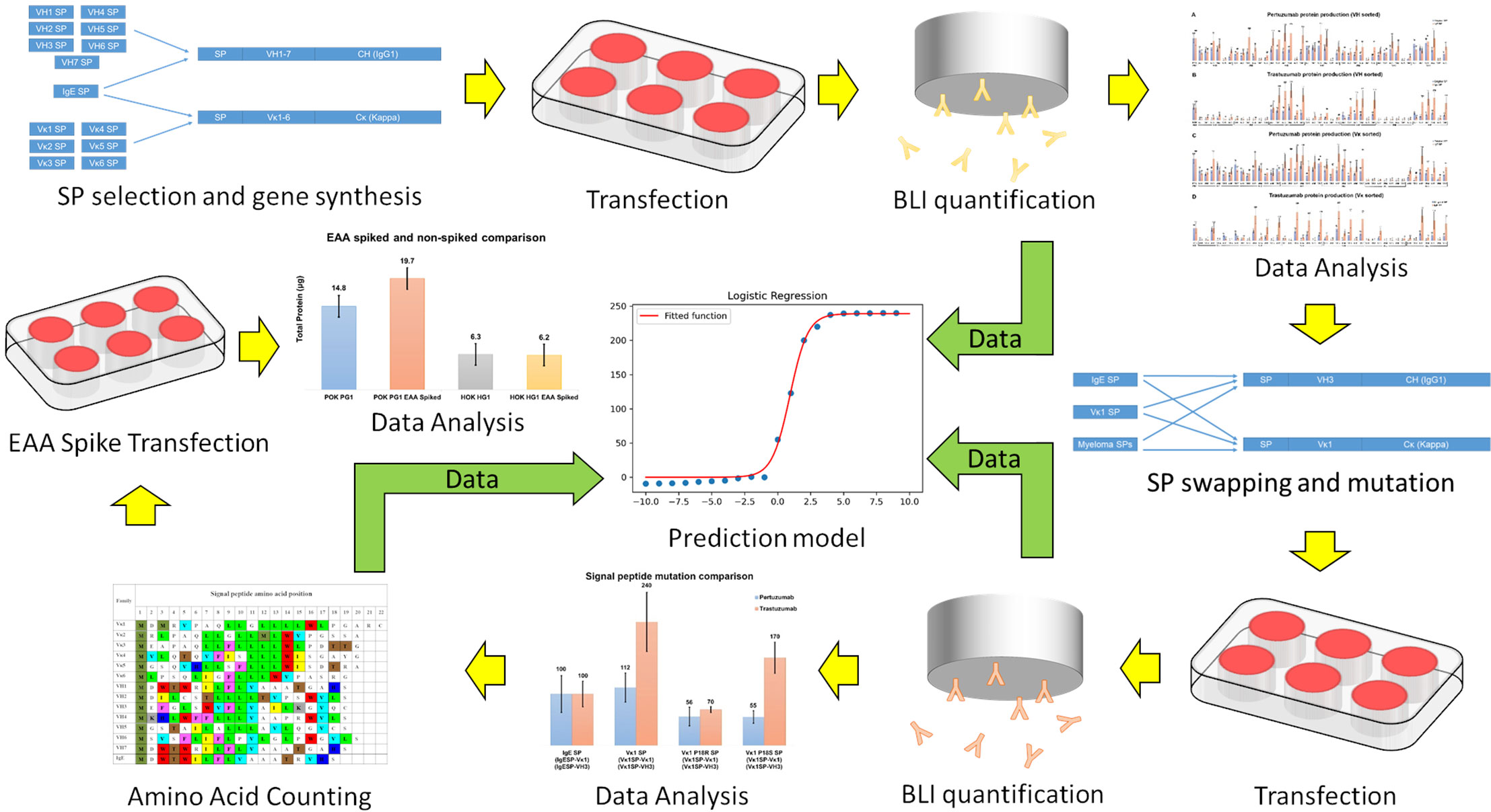
Frontiers | Essentially Leading Antibody Production: An Investigation of Amino Acids, Myeloma, and Natural V-Region Signal Peptides in Producing Pertuzumab and Trastuzumab Variants

The sensing of bacteria: emerging principles for the detection of signal sequences by formyl peptide receptors

Signal Peptide Optimization: Effect On Recombinant Monoclonal IgG Productivity, Product Quality And Antigen-Binding Affinity

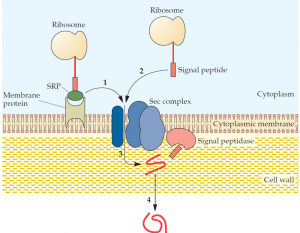
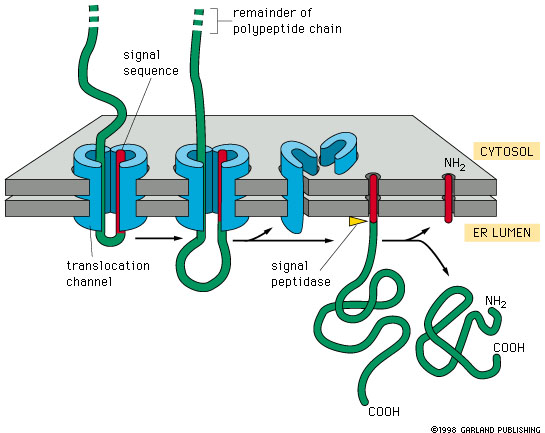
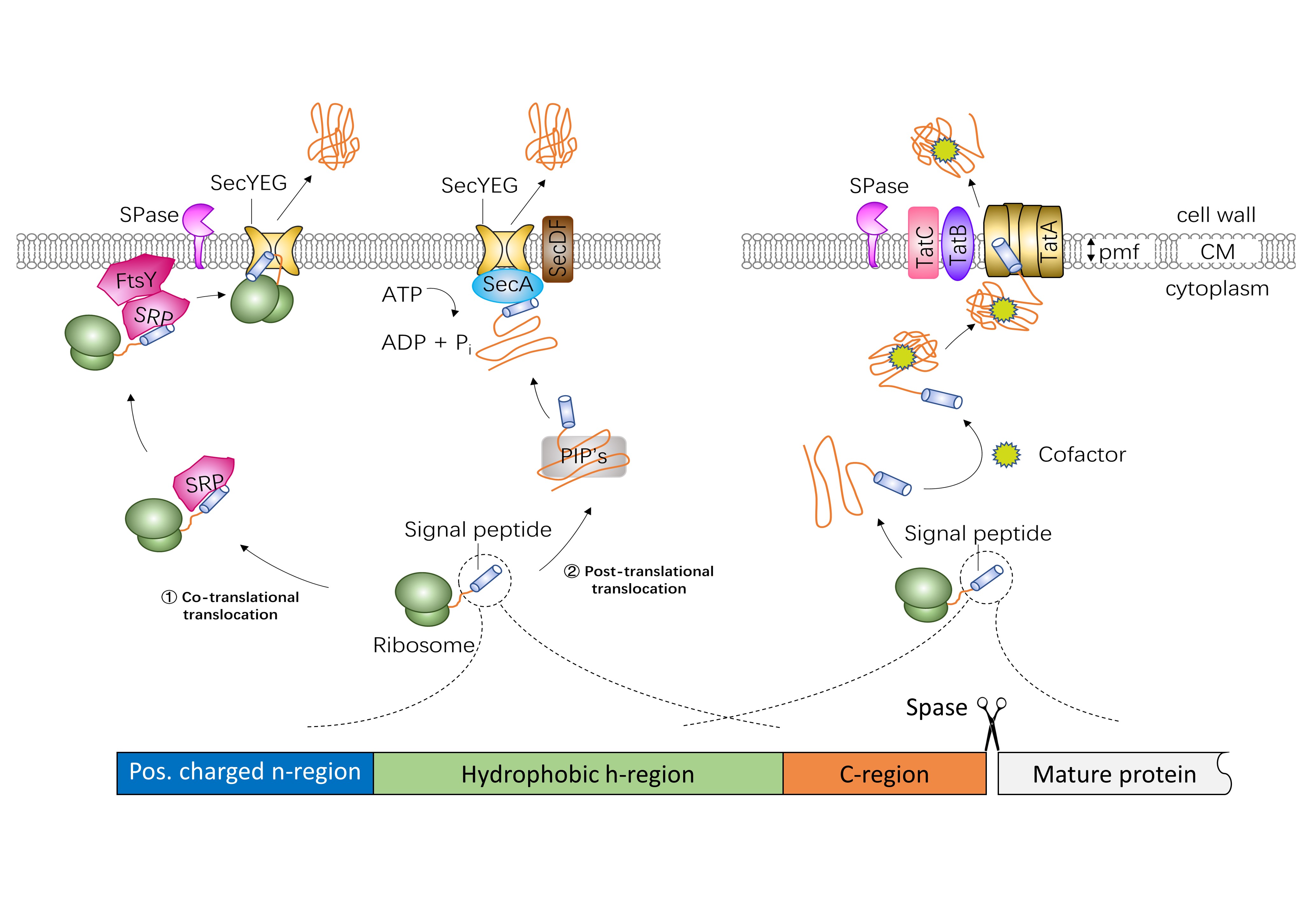
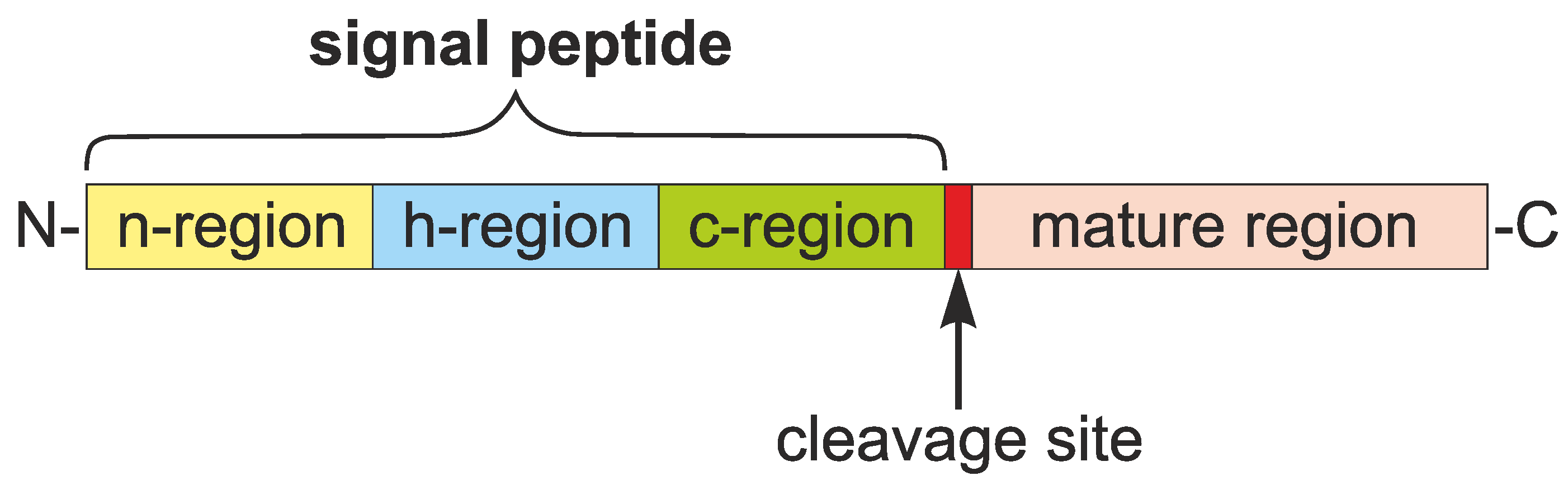
Diagram of the structure of the signal peptides and the translocation... | Download Scientific Diagram

Table 2 from Signal peptide design for improving recombinant protein secretion in the baculovirus expression vector system. | Semantic Scholar
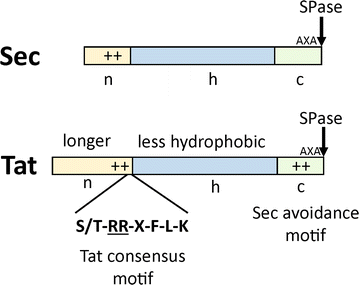
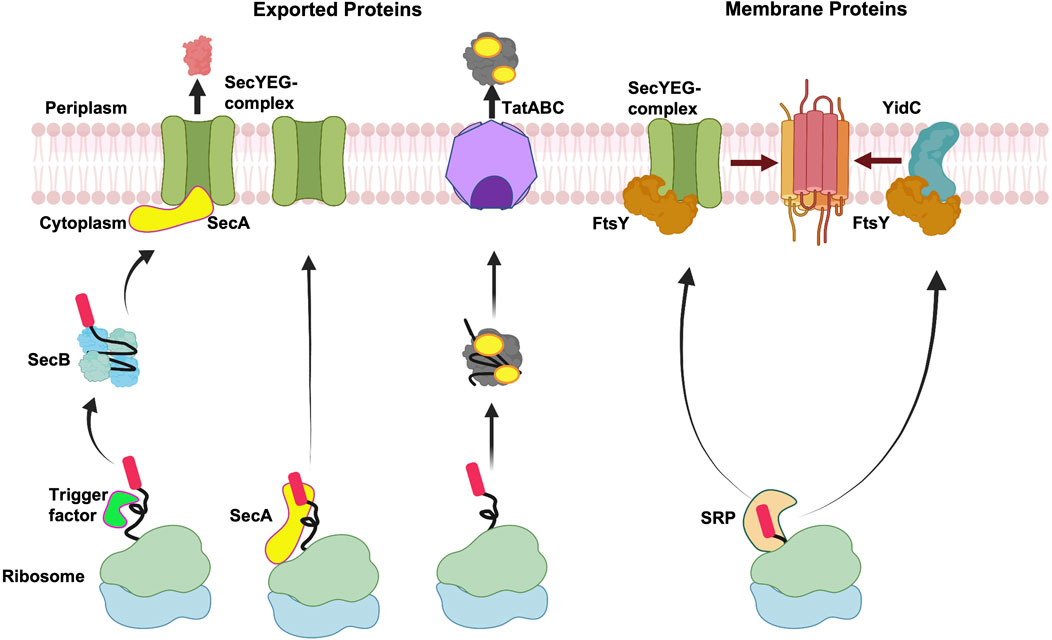
Signal peptides for recombinant protein secretion in bacterial expression systems | Microbial Cell Factories | Full Text

Cell re-entry assays do not support models of pathogen-independent translocation of AvrM and AVR3a effectors into plant cells | bioRxiv

Signal Peptide-Dependent Protein Transport inBacillus subtilis: a Genome-Based Survey of the Secretome | Microbiology and Molecular Biology Reviews

Synthetic pro‐peptide design to enhance the secretion of heterologous proteins by Saccharomyces cerevisiae - Cho - 2022 - MicrobiologyOpen - Wiley Online Library